Realm of truly big data
Humans are born with pattern recognition abilities, which enable us to discern patterns in graphic images at a glance. However, can we hope to visualize the relationship among the hundreds of variables in our massive data sets? Even the most advanced data visualization techniques do not go much beyond five dimensions.
Outline: Dimensional reduction process
High dimensional data
Dimensionality Reduction
Techniques for Dimensionality Reduction
Conclusion
Link to example exercise
What is high dimensional data?
In traditional dataset, the dimensions(p) of the data was low with many observations(n). In this case , classical rules such as the Central Limit Theorem are often applied to obtain some inference from data. A new challenge today is dealing with a different setting, when the data dimension(p) is very large and the number of observations(n) is small. So it means that whenever a dataset has p>n then it is high dimensional data.
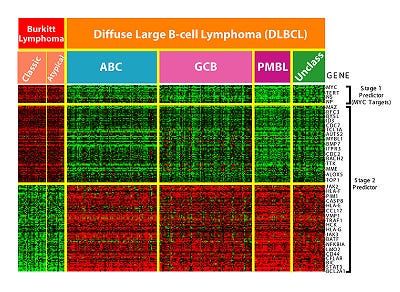
For example, there are few patients with many genes. In such cases, classical methods fail to produce a good understanding of the nature of the problem.
Issues with high dimensional data sets — Training a model on a dataset with many dimensions usually requires :
a) Vast time and space complexity
b) Often leads to overfitting
c) Not all feature are relevant to our problem
Thus we need a paradigm shift and a nonconventional way of thinking to problem-solving concerning high-dimensional data.
One most popular approach for mitigating high-dimensional data issues is dimensionality reduction.
2. What is dimensionality reduction all about?
Dimensionality reduction is the process of reducing the number of random variables under review, by obtaining a set of principal components.That is the data is converted from a high dimensional space into lower number of dimensions without loosing much of information.
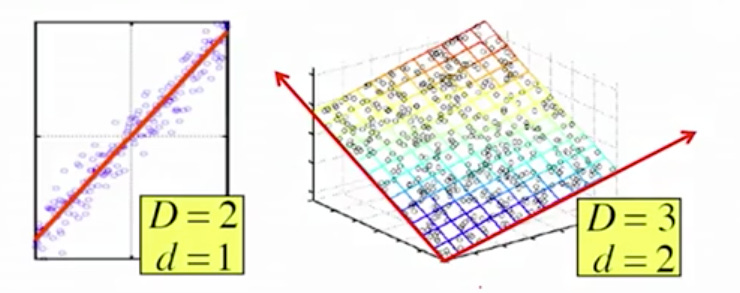
An example of dimensionality reduction can be discussed through a simple digital e-mail marketing classification problem, where we need to classify whether the e-mail is spam or not. This can involve a large number of features, such as whether or not the e-mail has a standard title, the content of the e-mail, whether the e-mail uses a personalised template, etc.Hence, we can reduce the number of features in such problems. A 3-D classification problem can be hard to visualize, whereas a 2-D one can be mapped to a simple 2 dimensional space, and a 1-D problem to a simple line.
The below figure illustrates this concept, where the idea is that these points are not randomly scattered in the space, they lie in the subset of the space. our goal is : to effectively find the subspace that represents the whole data.
Components of Dimensionality Reduction
There are two components of dimensionality reduction:
Feature selection: In this, we try to find a subset of the original set of variables, or features, to get a smaller subset which can be used to model the problem.
Feature extraction: This reduces the data in a high dimensional space to a lower dimension space, i.e. a space with lesser no. of dimensions.
3. How do we perform dimensionality reduction?
There are various methods used for dimensionality reduction but the most prevalent method in practice is : Principal Component Analysis (PCA)
Principal Component Analysis:
PCA is one of the most successful high dimensional analytics technique. This method was introduced by Karl Pearson. PCA seeks to explain the correlation structure of a set of predictor variables, using a smaller set of linear combinations of these variables. These linear combinations are called components.
It means that the total variability of a data set produced by the complete set of m variables can be accounted for by a smaller set of k linear combinations and there is almost as much information in the k components as there is in the original m variables.
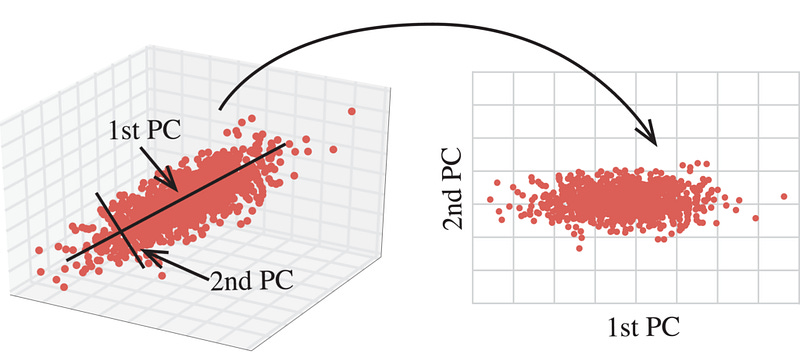
Suppose that the original variables form a coordinate system in m-dimensional space. The principal components represent a new coordinate system, found by rotating the original system along the directions of maximum variability.
We proceed to apply PCA using Eigen Decomposition or Singular Value Decomposition
a) Eigen decomposition involves the following steps:
Construct the covariance matrix of the data.
Compute the Eigen decomposition of the matrix — we get a set of Eigen values and Eigen vectors. I highly recommend you to watch this lecture Eigen vectors and Eigen values before you learn about the dimensionality reduction technique in detail, do not miss it!
Eigenvectors corresponding to the largest eigenvalues are used to reconstruct a large fraction of variance of the original data.
By doing so we get the following.
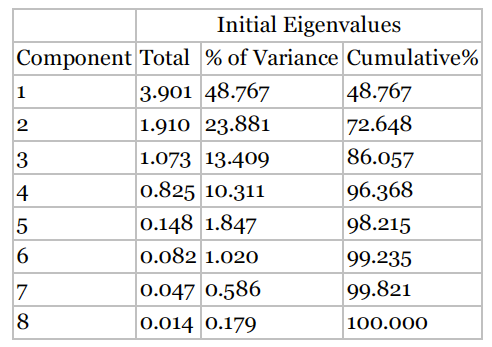
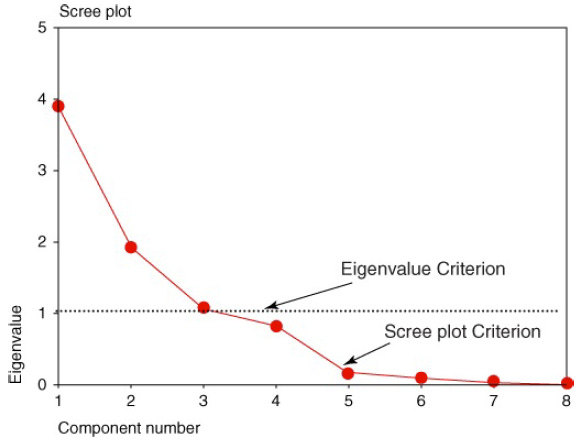
Recall that one of the motivations for PCA was to reduce the number of distinct explanatory elements. The question arises, “How do we determine how many components to extract?”
We can use scree plot to identify the number of components.A scree plot is a graphical plot of the eigenvalues against the component number. Scree plots are useful for finding an upper bound (maximum) for the number of components that should be retained.
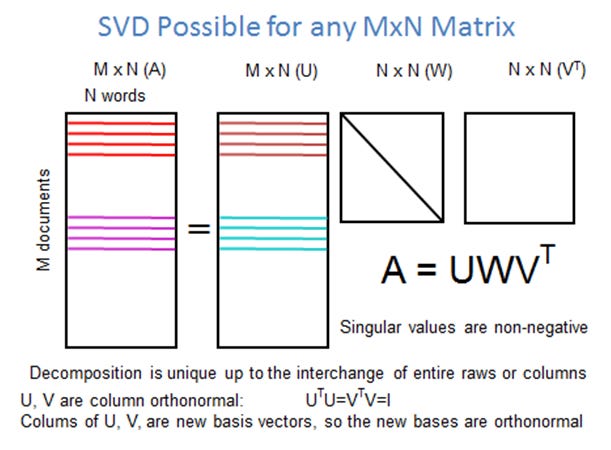
b) singular value decomposition:
The SVD is a direct method that generalises the concept of expressing a matrix as a diagonal matrix to any arbitrary matrix, as long as we use the right domain and range spaces.
Basic difference between Eigen decomposition and singular value decomposition is described below.
The Eigen decomposition uses only one basis, i.e. the eigenvectors, while he SVD uses two different basis, the left and right singular vectors
The basis of the Eigen decomposition is not necessarily orthogonal, the Eigen basis of the SVD is orthonormal!
It is proven that Singular Value Decomposition that is more efficient than the eigenvalue decomposition method depending on situations for analysis.To learn about the math behind the PCA analysis, watch the youtube lectures Part1,Part2 and Part3.
Conclusion:
Dimensionality reduction has proven useful in discovering non-linear, non-local relationships in the data that are not obvious in the feature space. In machine learning this is critical and hence powerful when applied.
PCA on IRIS dataset using Python:
Following are the steps to perform PCA and visualise the results using most famous IRIS dataset.
Dataset and Code:
https://github.com/ChitraRajasekaran/Principal-Component-Analysis
References: